Case report: First case of fungemia caused by Papiliotrema laurentii in a patient with SARS-CoV-2 infection
This is the first case report of fungemia and probable meningitis caused by Papiliotrema laurentii in a previously immunocompetent host with associated COVID-19
12/01/2024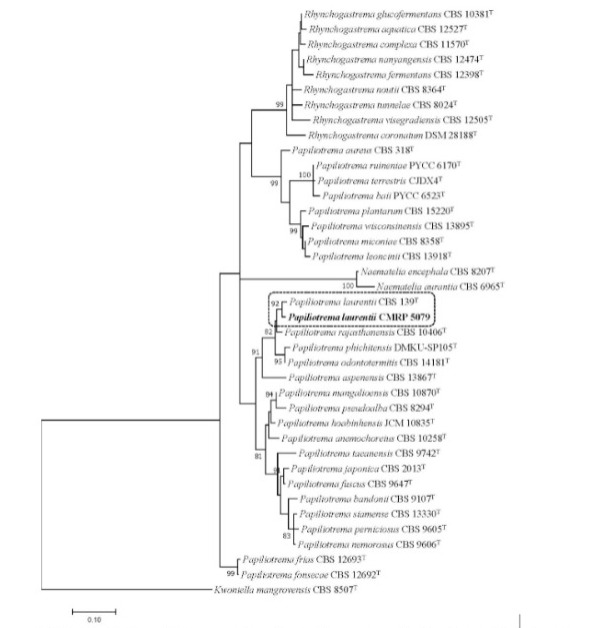
Phylogenetic tree was inferred using the maximum likelihood method based on the Tamura-Nei model with gamma distribution in the MEGA 7.0.26, and bootstrap support was calculated from 1000 replicates
Cognialli RCR et al. – Fungemia caused by Papiliotrema laurentii
Regielly Caroline Raimundo Cognialli[1], Gabriella Felber Fratucci[1], Bruno Hassunuma Carneiro[1], Karyna Leal Antonio[1], Morgana Ferreira Voidaleski[2], Larissa Molina Favarello[3], Vania Aparecida Vicente[4] and Flávio Queiroz-Telles[5]
[1]. Universidade Federal do Paraná, Hospital de Clínicas, Curitiba, PR, Brasil.
[2]. Universidade Federal do Paraná, Programa de Pós-Graduação em Microbiologia, Parasitologia e Patologia, Curitiba, PR, Brasil.
[3]. Universidade Federal de São Paulo, Escola Paulista de Medicina, São Paulo, SP, Brasil.
[4]. Universidade Federal do Paraná, Departamento de Patologia Básica, Curitiba, PR, Brasil.
[5]. Universidade Federal do Paraná, Hospital de Clínicas, Departamento de Saúde Coletiva, Curitiba, PR, Brasil.
Corresponding author: Dr. Flávio de Queiroz-Telles. e-mail: queiroz.telles@uol.com.br
Authors’ contribution
RCRC, GYF, BHC and FQ-T: Conceptualization;
RCRC, GYF, BHC and FQ-T: writing—original draft preparation;
RCRC, GYF, BHC, KLA, MFV, LMF, VAV and FQ-T: writing—review and editing;
RCRC, GYF, BHC, MFV, LMF, VAV and FQ-T: investigation and data curation.
Conflict of Interest
The authors declare no conflict of interest.
Orcid
Regielly Caroline Raimundo Cognialli: https://orcid.org/0000-0002-3895-1982
Gabriella Felber Fratucci: https://orcid.org/0009-0008-4013-7948
Karyna Leal Antonio: https://orcid.org/0000-0001-6299-1290
Bruno Hassunuma Carneiro: https://orcid.org/0000-0002-6022-2778
Morgana Ferreira Voidaleski: https://orcid.org/0000-0003-2751-352X
Larissa Molina Favarello: https://orcid.org/0000-0003-3739-9230
Vania Aparecida Vicente: https://orcid.org/0000-0002-2953-4861
Flávio de Queiroz-Telles: https://orcid.org/0000-0001-7034-2418
Abstract
Previously considered saprophytic and non-pathogenic, the fungus Papiliotrema laurentii(formerly known as Cryptococcus laurentii), is rarely associated with human infection. Nevertheless, there has been an increase in reported infections by non-neoformans cryptococci. After a literature search on the Cochrane Library, LILACS, SciELO, MEDLINE, PubMed, and PMC (PubMed Central) databases, we conclude that this is the first case report of fungemia and probable meningitis caused by Papiliotrema laurentii in a previously immunocompetent host with associated COVID-19.
Keywords: Papiliotrema, fungemia, SARS-CoV-2.
Introduction
Papiliotrema laurentii, formerly known as Cryptococcus laurentii, is rarely associated with human infection1. It can be found in the environment and has been previously considered a saprobe and non-pathogenic fungus2. However, in recent decades, the number of reported cases of infections caused by non-neoformans cryptococci, mainly Papiliotrema laurentii and Naganishia albida (formerly Cryptococcus albidus), has increased1-3. Fungemia caused by non-neoformans cryptococci has been more commonly reported in immunocompromised hosts, such as patients with HIV infection, lymphoproliferative disorders, malignancy, sarcoidosis, users of corticosteroid therapy, and after solid-organ transplantation2-5. Since the emergence of SARS-CoV-2, several reports have raised awareness of the association of COVID-19 with invasive fungal infections, including aspergillosis and candidiasis6.
A literature review was carried out in Cochrane Library, LILACS, SciELO, MEDLINE, PubMed, and PMC (PubMed Central) databases. We report the first case of fungemia caused by Papiliotrema laurentii in a previously immunocompetent patient with associated SARS-CoV-2 infection.
Case report
A 54-year-old Brazilian female patient, with a history of obesity, type 2 diabetes mellitus, heart failure, hypertension, and hypothyroidism presented to the emergency department with an 8-day history of cough, dyspnea, myalgia, fever, headache, and fatigue. She developed respiratory failure and underwent orotracheal intubation. A chest computed tomography scan (CT) revealed bilateral ground-glass opacities and consolidation in the left lower lobe. She received intravenous dexamethasone (6 mg/day), ceftriaxone, azithromycin, and prophylactic subcutaneous enoxaparin. SARS-CoV-2 was detected in a nasopharyngeal swab sample by reverse-transcription polymerase chain reaction (RT-PCR). The results of admission blood tests are summarized in Table 1.
Eleven days after admission, the patient developed severe acute kidney injury and shock; blood cultures were obtained, and antibiotics were changed. Initially, the regimen was changed to piperacillin-tazobactam and then, due to lack of improvement, escalated to meropenem plus vancomycin. After 72 h of incubation, the cultures were positive for yeasts, and micafungin (100 mg/day) was added to the anti-infective therapy. The yeasts were identified afterwards as Papiliotrema laurentii. The identification methods are summarized in Table 2. The species was confirmed as Papiliotrema laurentii by sequencing the large subunit (LSU) rRNA gene and internal transcribed spacer (ITS) region and clustered with P. laurentii CBS139, as shown in the phylogenetic analysis (Figure 1)7. The isolate was deposited in the Microbiological Collections of the Paraná Network – Taxonline (CMRP) as Papiliotrema laurentii CMRP 5079. Antifungal susceptibility tests were performed using the microbroth dilution method according to the Clinical and Laboratory Standards Institute (CLSI) (Table 2). A total of 15 blood cultures were collected during hospitalization. Papiliotrema laurentii was isolated from two independent blood cultures, with a difference of 6 days between each other (on days 11 and 17). At the time of the collection of the first positive blood culture, the patient had leukocytosis (18.980 cells/mm³) and neutrophilia (17.272 cells/mm³). In order to rule out central nervous system involvement, head CT and lumbar puncture were performed. Imaging was unremarkable, and cerebrospinal fluid (CSF) analysis revealed pleocytosis (9 cells/mm³) with lymphocyte predominance (88%), hyperproteinorrachia (377 mg/dL), and glucose and lactate levels of 107 mg/dL and 3.38 mmol/L, respectively. The CSF opening pressure was normal, no microorganisms were seen on direct microscopy, and cultures were negative. Despite negative microbiological results in the initial CSF analysis, induction therapy for severe-disseminated disease was initiated based on the presence of fungemia and lymphocytic meningitis with fluconazole (FLC) (800 mg/day) plus amphotericin B (AmpB) deoxycholate (50 mg/day). Serial blood cultures and CSF samples were drawn for treatment monitoring; follow-up blood cultures were negative, CSF leukocyte count normalized (0 cell/mm³) after 1 week, and CSF protein levels reduced to 77 mg/dL after 2 weeks of antifungal therapy. Induction therapy with FLC plus AmpB was prescribed for 2 weeks, and consolidation therapy with FLC (800 mg/day) was maintained for 2 months afterwards. FLC (300 mg/day) was indicated for maintenance therapy, and the patient was discharged with oxygen support via tracheostomy after a 3-month hospitalization.
Discussion
Cryptococcal infections are caused by encapsulated yeast-like fungi of the genus Cryptococcus; the most common species complexes are C. neoformans and C. gattii. These microorganisms are typically found on soil contaminated with pigeon excreta and trees. Infections caused by P. laurentii are relatively uncommon; they are more frequently described in immunocompromised patients2-5. Risk factors for infection include impairment of cell-mediated immunity, HIV/AIDS, hematologic malignancies, neutropenia, corticosteroid treatment, chemotherapy, solid-organ transplantation, impaired humoral immunity, non-HIV-related lymphopenia, and direct or indirect exposure to pigeon excreta. Clinical manifestations include skin and ocular lesions, lung infection, peritonitis, and meningitis. Patients can also present with systemic symptoms such as fever and septic shock. Most fungemia cases have been reported in previously immunocompromised patients5.
SARS-CoV-2 infection can be associated with different levels of immunosuppression. The immunosuppression induced by moderate-to-severe COVID-19 is aggravated by exposure to corticosteroids and invasive devices, which may predispose individuals to infection by uncommon pathogens, such as P. laurentii6,8. Cryptococcemia has been associated with COVID-19 during the use of immunosuppressive agents and corticosteroids8,9. Other fungal infections have been described in COVID-19 patients, including pulmonary aspergillosis, invasive candidiasis, and mucormycosis6,9. In 2022, a case of endogenous endophthalmitis caused by P. laurentii was described in a post-COVID-19 patient who had received corticosteroid therapy10.
Evidence guiding the treatment of non-neoformans cryptococcal infections is limited by the paucity of cases and the lack of clinical studies. For P. laurentii bloodstream infection, AmpB has been used in several cases, mostly in association with flucytosine and FLC2. Monotherapy with FLC has also been successful in some patients with fungemia3. Similarly to fungemia, data available on the management of meningitis are limited to case reports. The reported duration of therapy ranges from 14 days in patients with fungemia to 36-150 days in patients with meningitis and endophtalmitis10. Nevertheless, the treatment strategies used in this case were in accordance with clinical practice guidelines for the management of severe-disseminated cryptococcal disease11. Although P. laurentii CMRP 5079 showed reasonably high FLC minimal inhibitory concentrations (MIC) in vitro (4-8 μg/mL), resistance in vivo was not observed, as the patient had favorable clinical and microbiological outcomes. It is important to note that there are currently no interpretative breakpoints available for Cryptococcus/Papiliotrema spp. regarding FLC resistance12.
In conclusion, the present study describes the first case of fungemia and probable meningitis caused by P. laurentii in a previously immunocompetent patient with COVID-19. Considering the vulnerability of patients with severe COVID-19 to coinfections, early suspicion and detection of fungal pathogens are necessary to reduce morbidity and mortality.
Acknowledgments
We thank the team of the Infectious Diseases Department and Laboratory of Hospital de Clínicas from Federal University of Paraná.
Vania Aparecida Vicente received fellowships from National Council for Scientific and Technological Development (CNPq), Brasilia, Brazil (http://cnpq.br/), grant number 312811/2018-7, and grants support from the CNPq project number 402509/2022-6.
References
- Banerjee P, Haider M, Trehan V, Mishra B, Thakur A, Dogra V, et al. Cryptococcus laurentii fungemia. Indian J Med Microbiol. 2013;31(1):75-7.
- Londero MR, Zanrosso CD, Corso LL, Michelin L, Soldera J. Catheter-related infection due to Papiliotrema laurentii in an oncologic patient: Case report and systematic review. Braz J Infect Dis. 2019;23(6):451-61.
- Neves RP, Lima Neto RG, Leite MC, Silva VK, Santos Fde A, Macêdo DP. Cryptococcus laurentii fungaemia in a cervical cancer patient. Braz J Infect Dis. 2015;19(6):660-3.
- Intra J, Sarto C, Brambilla P. A rare case of cutaneous Papiliotrema (Cryptococcus) laurentii infection in a 23-year-old Caucasian woman affected by an autoimmune thyroid disorder with hypothyroidism. Eur J Clin Microbiol Infect Dis. 2021;40(3):647-50.
- Al-Otaibi H, Asadzadeh M, Ahmad S, Al-Sweih N, Joseph L. Papiliotrema laurentii fungemia in a premature, very low-birth-weight neonate in Kuwait successfully treated with liposomal amphotericin B. J Mycol Med. 2021;31(2):101123.
- Kangabam N, Nethravathy V. An overview of opportunistic fungal infections associated with COVID-19. 3 Biotech. 2023;13(7):231.
- Kurtzman CP, Robnett CJ. Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek. 1998;73(4):331-71.
- Khatib MY, Ahmed AA, Shaat SB, Mohamed AS, Nashwan AJ. Cryptococcemia in a patient with COVID-19: A case report. Clin Case Rep. 2020;9(2):853-5.
- Abdoli A, Falahi S, Kenarkoohi A. COVID-19-associated opportunistic infections: a snapshot on the current reports. Clin Exp Med. 2022;22(3):327-46.
- Deepa MJ, Megharaj C, Patil S, Rani PK. Cryptococcus laurentii endogenous endophthalmitis post COVID-19 infection. BMJ Case Rep. 2022;15(5):e246637.
- Mourad A, Perfect JR. Present and Future Therapy of Cryptococcus Infections. J Fungi (Basel). 2018;4(3):79.
- Espinel-Ingroff A, Cantón E. Methods for Antifungal Susceptibility Testing of the Cryptococcus neoformans/C. gattii Complex: Strengths and Limitations. J Fungi (Basel). 2023;9(5):542.
TABLE 1: Laboratory results from admission blood tests.
| Blood tests | Results |
| Leukocyte count (cells/mm³) | 11,860 |
| Lymphocytes (cells/mm³) | 593 |
| Platelets (cells/mm³) | 265,000 |
| D-Dimer level (mg/L) | 1.8 |
| Fibrinogen level (mg/dL) | 485 |
| Lactate dehydrogenase level (U/L) | 778 |
TABLE 2: Identification of Papiliotrema laurentii CMRP 5079 and antifungal susceptibility tests.
| Yeast identification | Results | |||||||||
| Time of incubation for growth in BACTEC Plus Aerobic/FBD culture medium (Becton-Dickinson, Sparks, USA) | 72h | |||||||||
| Colonies on blood agar | Smooth and cream colored | |||||||||
| Nigrosin stain | Round-to-oval yeasts with slight capsule | |||||||||
| Temperature of growth | 30 °C optimum growth
37 °C poor growth |
|||||||||
| Urease | Positive | |||||||||
| Niger media | No melanin pigment | |||||||||
| Vitek2 (bioMerieux, Durham, NC) | Cryptococcus laurentii | |||||||||
| MALDI-TOF Vitek MS (bioMerieux, Durham, NC) | Cryptococcus laurentii | |||||||||
| Cryptococcal antigen lateral flow assay in blood (CrAg LFA; IMMY Inc., Norman, OK) | Negative | |||||||||
| Cryptococcal antigenlatex agglutination Pastorex Crypto Plus in blood (Bio-Rad, Marnes-la-Coquette, France) | Negative | |||||||||
| LSU rRNA and ITS region sequencing | Papiliotrema laurentii | |||||||||
| Antifungal susceptibility tests for Papiliotrema laurentiiCMRP 5079 isolates | ||||||||||
| Amphotericin B | Fluconazole | Voriconazole | ||||||||
| 24h | 48h | 72h | 24h | 48h | 72h | 72h | 24h | 48h | ||
| P. laurentiiCMRP 5079 I (MIC, μg/mL) | 0.25 | 0.25 | 0.5 | 4 | 8 | 8 | 0.125 | 0.125 | 0.25 | |
| P. laurentiiCMRP 5079 II (MIC, μg/mL) | 0.25 | 0.25 | 0.5 | 4 | 8 | 8 | 0.125 | 0.125 | 0.25 | |
MIC = Minimal Inhibitory Concentration; LSU rRNA= Large subunit ribosomal ribonucleic acid; ITS = Internal Transcribed Spacer

FIGURE 1: Phylogenetic tree was inferred using the maximum likelihood method based on the Tamura-Nei model with gamma distribution in the MEGA 7.0.26, and bootstrap support was calculated from 1000 replicates. Bootstrap values ≥ 80% were considered statistically significant. The sequencer of ITS region and the D1/D2 region of the LSU rRNA gene showed the isolate CMRP 5079 was close to Papiliotrema laurentii CBS 139 type strain. Kwoniella mangrovensis CBS 8507T was taken as outgroup. (T) = type strain of the species or of one of its synonyms.